MirrorTree
Predictor of protein interaction partners and functional relationships based on similarity of phylogenetic histories.
References
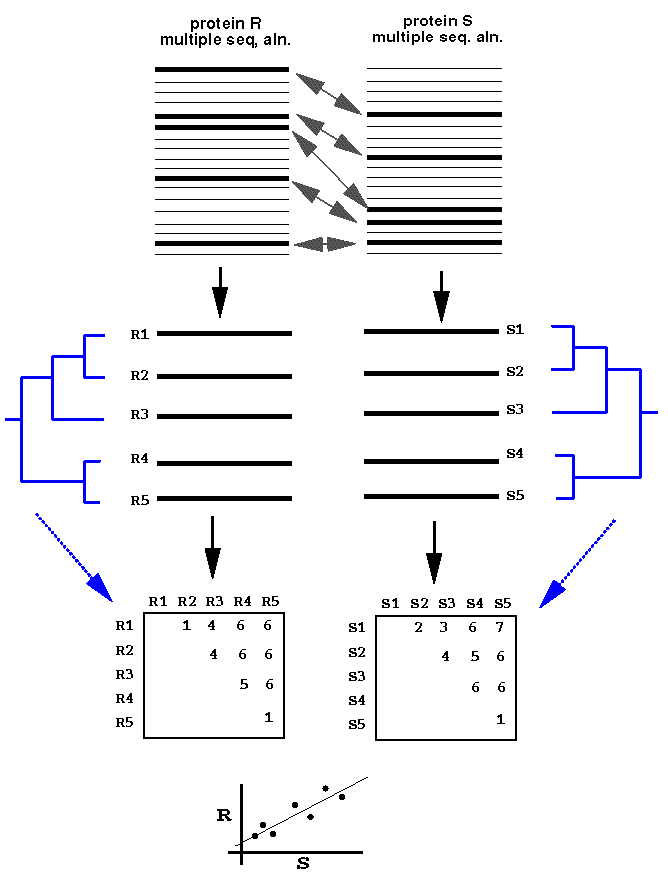
- Florencio Pazos & Alfonso Valencia (2001).
Similarity of phylogenetic trees as indicator of protein-protein interaction.
Protein Engineering. 14(9):609-614.
[MedLine:11707606]
[HTML]
[PDF]
- Florencio Pazos, Juan A. G. Ranea, David Juan & Michael J. E. Sternberg. (2005).
Assessing Protein Co-evolution in the Context of the Tree of Life Assists in the Prediction of the Interactome.
Journal of Molecular Biology.
352(4):1002-1015.
[PubMed:16139301]
[HTML]
[PDF]
[online additional material]
- David Juan, Florencio Pazos & Alfonso Valencia. (2008).
High-confidence prediction of global interactomes based on genome-wide coevolutionary networks.
Proc Natl Acad Sci USA.
105(3):934-939.
[PubMed:18199838]
[HTML]
[PDF]
[ECID system]
- David Ochoa & Florencio Pazos. (2010).
Studying the co-evolution of protein families with the Mirrortree web server.
Bioinformatics.
26(10):1370-1371.
[Pubmed:20363731]
[HTML]
[PDF]
[MirrorTree web server]
-
Dorota Herman, David Ochoa, David Juan, Daniel Lopez, Alfonso Valencia and Florencio Pazos (2011).
Selection of organisms for the co-evolution-based study of protein interactions.
BMC Bioinformatics.
12:363.
[PubMed:21910884]
[HTML]
[PDF]
-
David Ochoa, Ponciano García-Gutiérrez, David Juan, Alfonso Valencia and Florencio Pazos. (2013).
Incorporating information on predicted solvent accessibility to the co-evolution-based study of protein interactions.
Molecular Biosystems.
9(1):70-76.
[PubMed:23104128]
[HTML]
[PDF]
[Mirrortree web server]
- David Ochoa, David Juan, Valencia, A. and Florencio Pazos (2015).
Detection of significant protein co-evolution.
Bioinformatics.
31(13):2166-2173.
[PubMed:25717190]
[HTML]
[PDF]
[pMT software]
[E coli co-evolutionary network]
[Mirrortree web server]
Recent reviews
Datasets
- Datasets used in these publications are available under request.
Software
Which software/mirrortree variant to use? From the following list, choose the method closer to the top depending on the data and bioinformatics expertise you have.
- If you have a genome-wide collection of multiple alignments/phylogenetic trees and some bioinformatics expertise, use the
pMT methodology (Ochoa et al. 2015), or alternatively ProfileCorrelation/ContextMirror (Juan et al., 2008).
- If you only have a small set of alignments/trees (in the limit only those for your two proteins of interest) but you have a tree with inter-species distances (e.g. that based on the 16SrRNA) and some expertise in bioinformatics, use the tol-mirrortree approach (Pazos et al. 2005).
- If you only have a small set of alignments/trees (in the limit only those for your two proteins of interest) and some expertise in bioinformatics, use the original mirrortree approach (Pazos et al. 2001) and pay attention to the composition of the alignments (in terms of sequence redundancy, etc.) as explained in Herman et al. (2011).
- If you do not have neither alignments/trees, nor bioinformatics expertise, use the mirrortree web server which requires single sequences of your two proteins of interest to run.
Dr. Florencio Pazos Cabaleiro.
Computational Systems Biology Group.
National Center for Biotechnology (CNB-CSIC)
C/Darwin, 3. Cantoblanco. 28049 Madrid. Spain
Tlf. +34.915854669. Fax. +34.915854506