Tools for the functional interpretation of metabolomic experiments
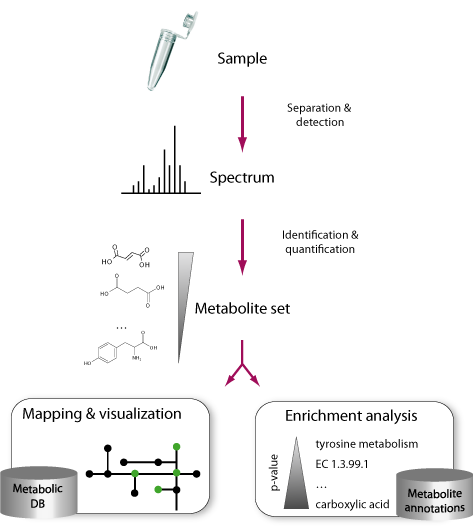
Omics approaches (e.g genomics, transcriptomics, proteomics) seek to massively characterizing the molecular repertories of living systems. Among them, metabolomics tries to determine the set of metabolites in a given biological system. As metabolomic techniques develop and allow characterizing larger sets of metabolites, researchers need automatic methods to interpret their experimental results.
We can classify these tools in two groups:
More information and links
Servers and tools
| Name | URL |
|---|---|
| BioCyc - Omics Viewer | http://biocyc.org |
| iPath | http://pathways.embl.de |
| KaPPA-View | http://kpv.kazusa.or.jp/en/ |
| KEGG | http://www.genome.jp/kegg/pathway.html |
| MapMan | http://mapman.gabipd.org/web/guest/mapman |
| MetPA | http://metpa.metabolomics.ca |
| Metscape | http://metscape.ncibi.org |
| MGV | http://www.microarray-analysis.org/mayday |
| Paintomics | http://www.paintomics.org |
| Pathos | http://motif.gla.ac.uk/Pathos/ |
| Pathvisio | http://www.pathvisio.org/ |
| ProMetra | http://www.cebitec.uni-bielefeld.de/groups/brf/software/prometra_info/ |
| Reactome | http://www.reactome.org |
| VANTED | http://vanted.ipk-gatersleben.de |
| Name | URL |
|---|---|
| IMPaLA | http://impala.molgen.mpg.de |
| MBROLE 2.0 | http://csbg.cnb.csic.es/mbrole2 |
| MPEA | http://ekhidna.biocenter.helsinki.fi/poxo/mpea/ |
| MSEA | http://www.msea.ca |
© 2012, Computational Systems Biology Group. CNB-CSIC