 |
||
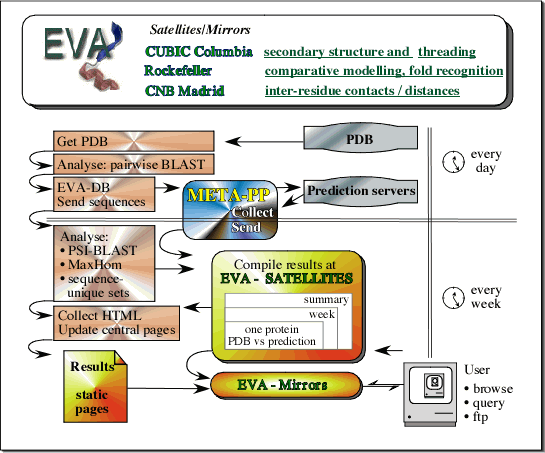
Flowchart of EVA. Every day, EVA downloads the newest protein structures from PDB [1] . The structures are added to mySQL databases, sequences are extracted for every protein chain, and are sent to each prediction server by META-PredictProtein [2] . META-PP collects the results and sends them to EVA. Every week, EVA runs alignment programs for searching sequence (iterated PSI-BLAST [3] , MaxHom [4] ) and structure (CE [5] , ProSub [6] ) databases to determine homologues. Predictions of secondary structure and inter-residue contacts, as well as, comparative modelling are evaluated at the EVA satellites at Columbia University, Rockefeller University, and CNB Madrid. The central EVA site at Columbia collects all the assessments from the satellites and the results from the database searches, and publishes the updated web pages. Finally, all web pages are mirrored at the satellites.